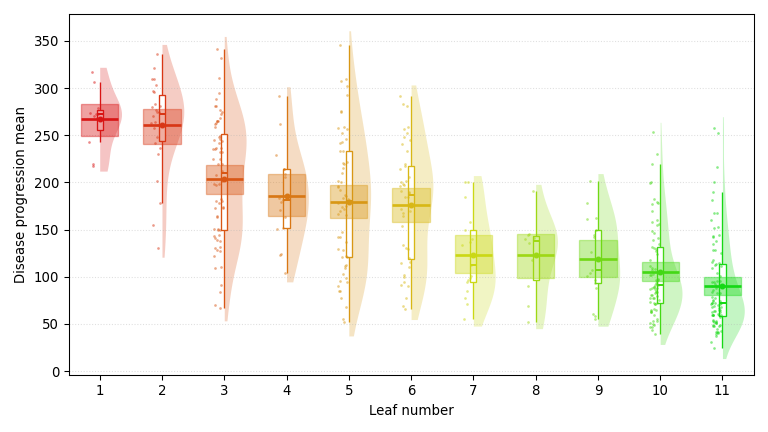Sigma
- Conditional inference trees library in Python, designed for quick, robust and easy diagnostics of data in production
- Fully transparent and statistically rigorous, with significance tests and confidence intervals at every step
- Source-available, production-ready with minimal dependencies and a simple API
Source-Available
Sigma is freely available under a source-available license. Contributions, issues and feedback are welcome.
View on GitHubSee It in Action
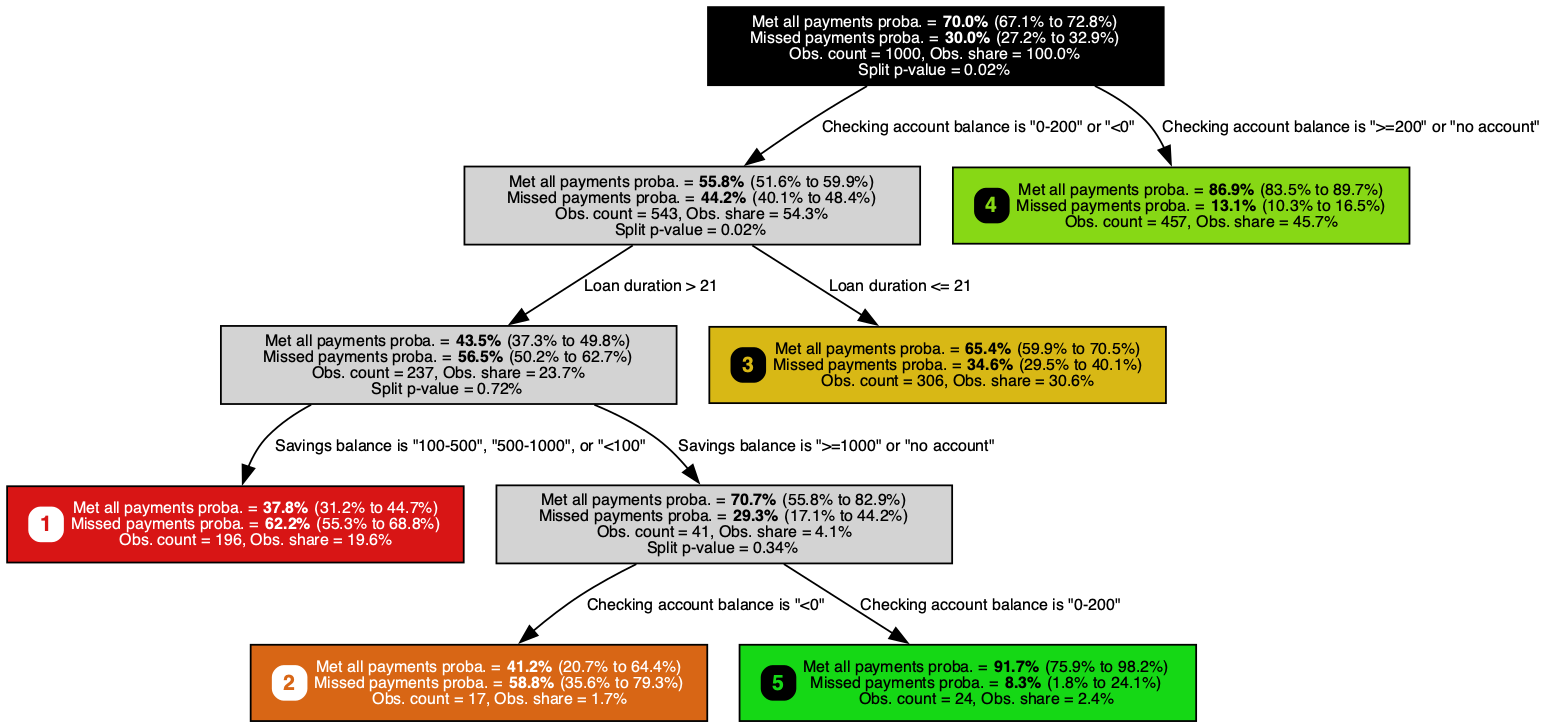
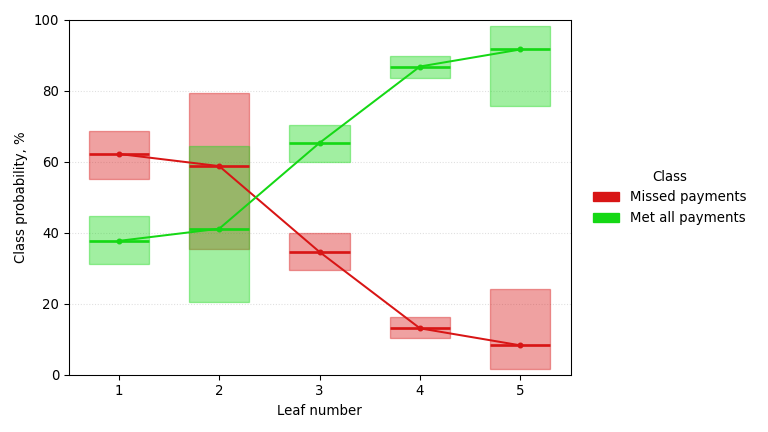
A classification tree trained on the German Credit dataset, predicting missed-payment probability (smaller is better) with a Jeffreys 95% confidence interval at each node. It pulls out checking account balance, loan duration, and savings balance, recovering the signals retail banks have long weighed alongside credit-bureau data.
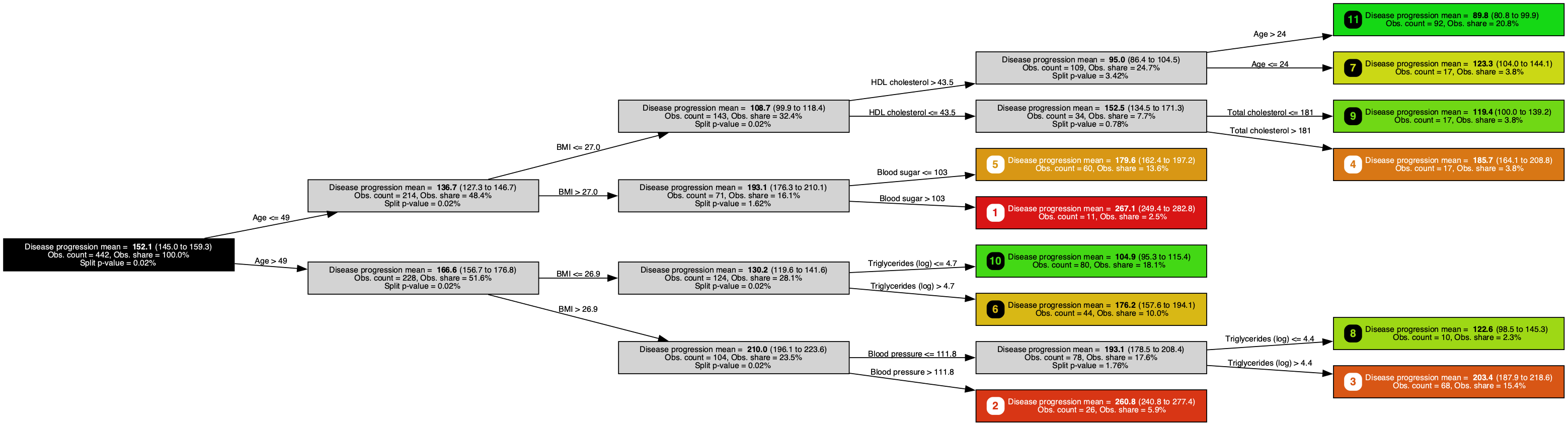
A regression tree trained on the Diabetes dataset, predicting one-year disease progression (smaller is better) with a Bayesian-bootstrap 95% confidence interval at each node. It surfaces age, BMI, and HDL cholesterol, echoing the demographic and lipid-panel staples behind every diabetes-risk calculator.
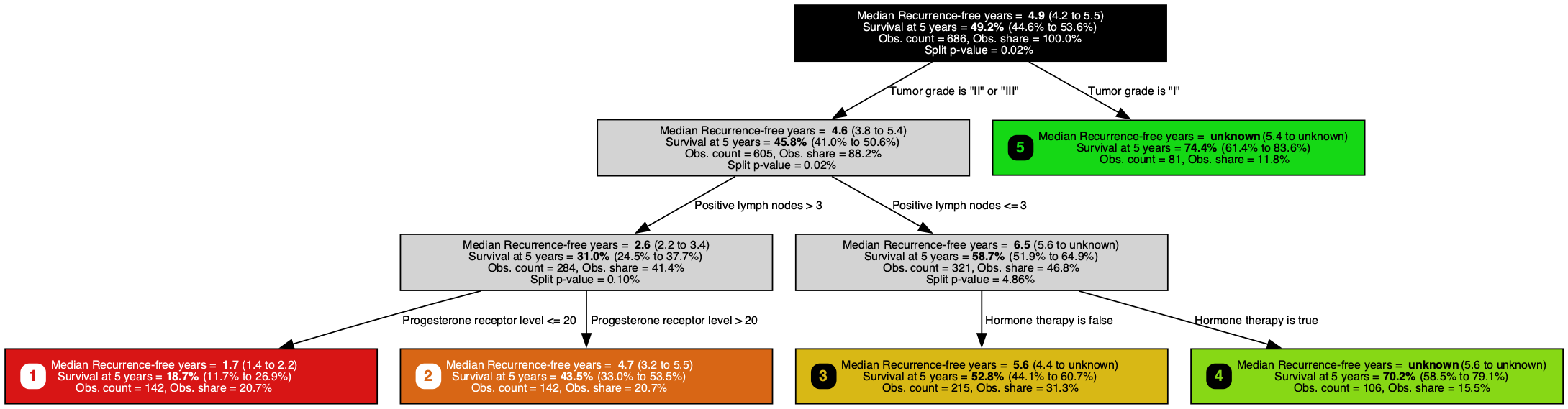
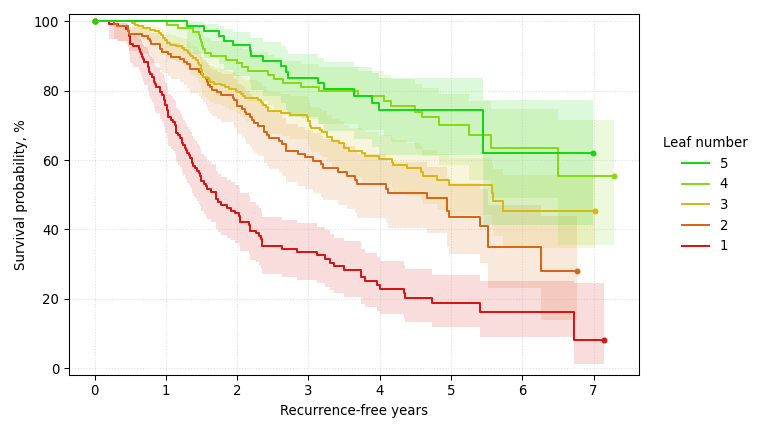
A survival tree trained on the GBSG-2 breast cancer dataset, predicting recurrence-free days (larger is better) with a Brookmeyer-Crowley 95% confidence interval at each node. It splits on positive lymph nodes, hormone therapy, and progesterone receptor level, the same staging, treatment, and biomarker variables clinicians already record on every breast cancer chart.
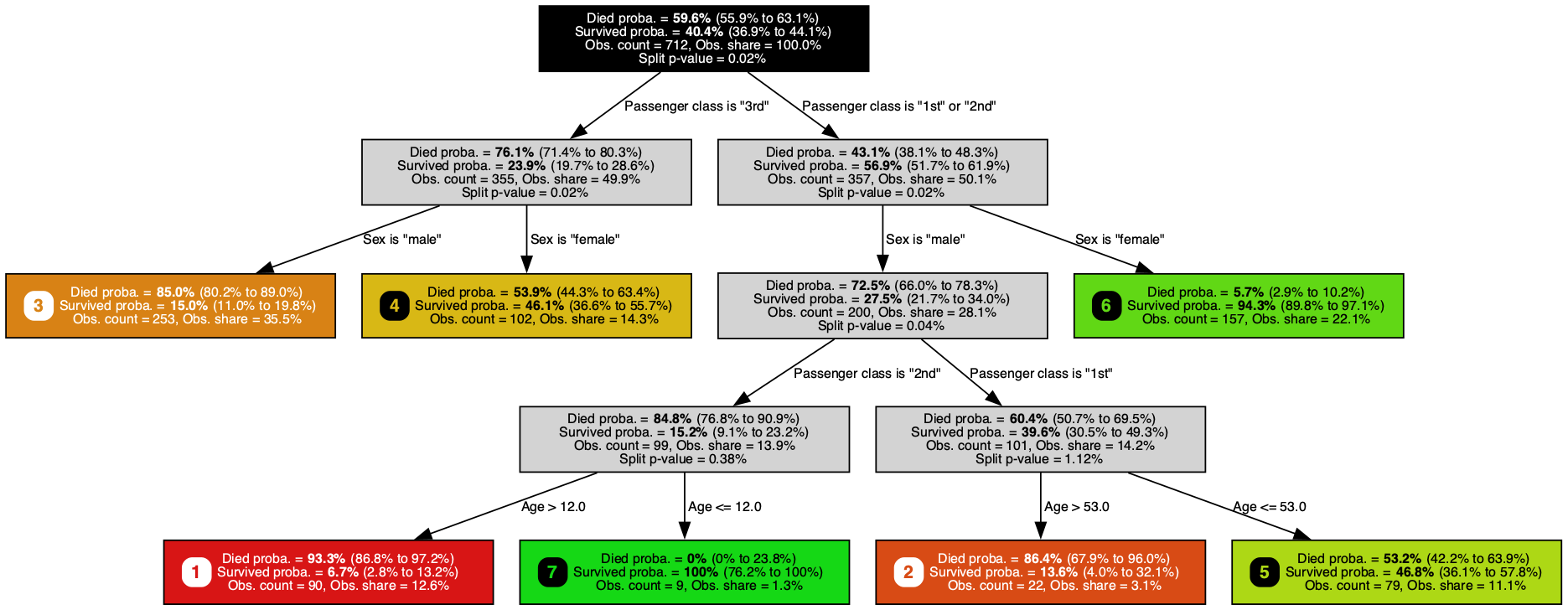
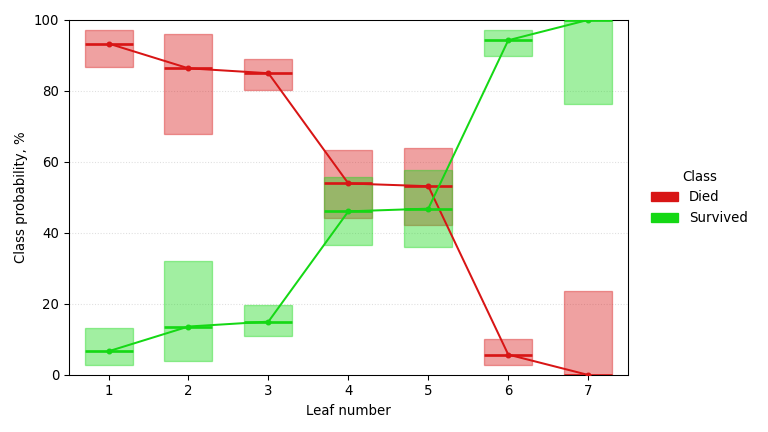
A classification tree trained on the Titanic dataset, predicting survival probability (larger is better) with a Jeffreys 95% confidence interval at each node. It singles out passenger class, sex, and age, rediscovering the women-and-children-first dynamics of that night from nothing but a passenger manifest.
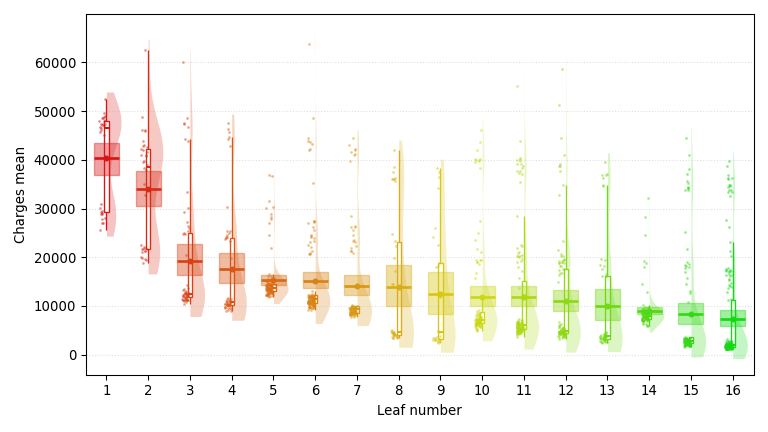
A regression tree trained on the Insurance dataset, predicting medical insurance charges (smaller is better) with a Bayesian-bootstrap 95% confidence interval at each node. It highlights age, smoking status, and number of children, three of the core demographic and behavioral factors health insurers build premiums around.
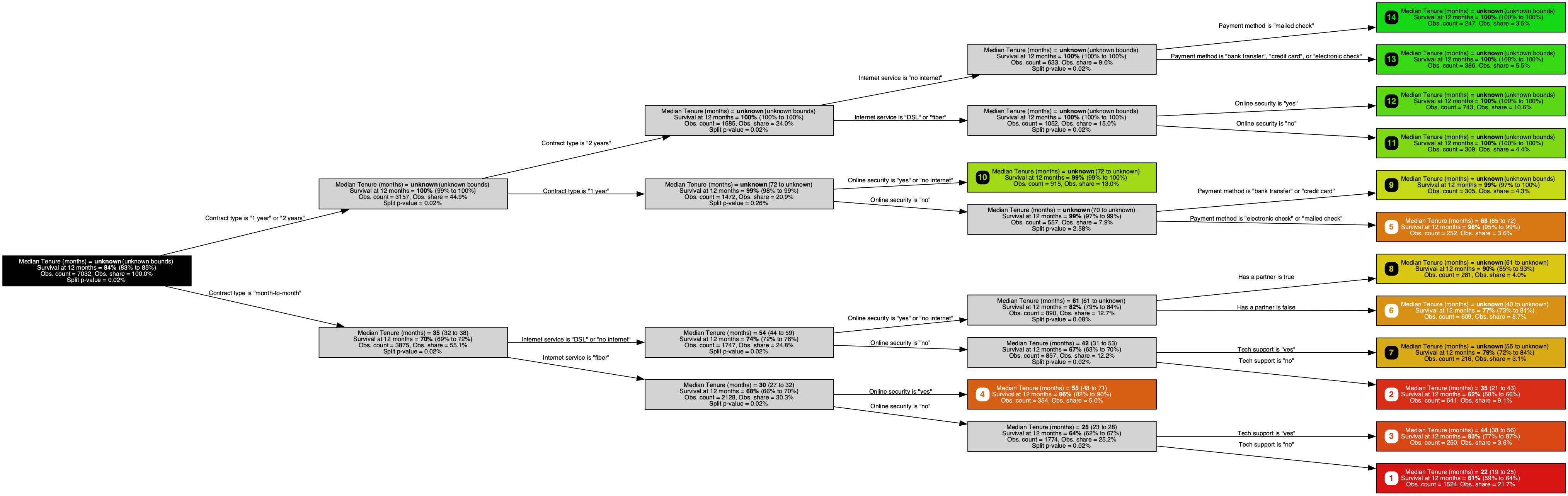
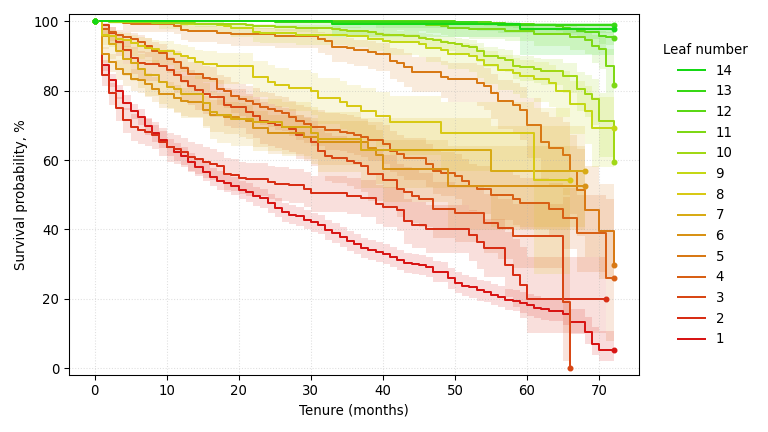
A survival tree trained on the IBM Telco Customer Churn dataset, predicting time to churn (larger is better) with a Brookmeyer-Crowley 95% confidence interval at each node. It homes in on contract type, internet service, and online security, three of the contract and service-tier signals subscription analysts already weigh when modeling churn.
Key Features
Instant Diagnostic
On a fresh dataset, one fit is often enough to surface the prediction potential, key predictors, leading interactions, and obvious data-quality issues. This avoids committing time to a full modeling effort. All feature types are supported, so an initial diagnostic needs no upstream preprocessing pipeline.
When a production model starts drifting or failing to pick up a new behavior, Sigma quickly reveals where and how the data or model is shifting, without diverting significant data-scientist time.
Point Sigma at an existing model to compare its predictions against observed outcomes. The resulting tree pinpoints exactly where, why and how the model fails.
Full Transparency
Sigma's model is a tree, not a black box: a few compact, readable rules, inspectable end-to-end.
Splits use significance tests (Hothorn, Hornik, and Zeileis, 2006) with multiplicity correction, so signals are separated from noise on statistical grounds. Tree branches are statistically validated, and growth stops once no signal remains. CART's bias toward features with many categories is also removed.
Leaf predictions come with confidence intervals. Several variants (e.g., Jeffreys, Clopper-Pearson, Bayesian bootstrap, Student-t) let you pick the coverage guarantee you need, so uncertainty stays explicit at every leaf.
Production-Ready
Sigma installs with
pip install ars-sigma
and depends only on the industry-standard NumPy, SciPy, and
scikit-learn. It runs on Python 3.10+ with no compiled
extensions to build.
RegressionTree,
ClassificationTree, and
SurvivalTree
implement the scikit-learn API, fitting into your existing
pipelines, cross-validation, and serialization without extra
wiring. Pandas DataFrames are supported throughout.
Sample weights are supported end-to-end, and fits are
reproducible from a fixed
random_state.
Trained models are plain Python state that pickles cleanly
and moves from a training notebook to a batch job or
serverless runtime with no platform-specific build step.
Grounded in Research
Every statistical method in Sigma comes from a peer-reviewed paper.
Hothorn, T., & Zeileis, A. (2015). partykit: A Modular Toolkit for Recursive Partytioning in R. Journal of Machine Learning Research, 16, 3905-3909. jmlr.org/papers/v16/hothorn15a
Patil, V. V., & Kulkarni, H. V. (2012). Comparison of Confidence Intervals for the Poisson Mean: Some New Aspects. REVSTAT - Statistical Journal, 10(2), 211-227. doi:10.57805/revstat.v10i2.117
Hothorn, T., Hornik, K., & Zeileis, A. (2006). Unbiased Recursive Partitioning: A Conditional Inference Framework. Journal of Computational and Graphical Statistics, 15(3), 651-674. doi:10.1198/106186006X133933
Hothorn, T., Hornik, K., van de Wiel, M. A., & Zeileis, A. (2006). A Lego System for Conditional Inference. The American Statistician, 60(3), 257-263. doi:10.1198/000313006X118430
Olsson, U. (2005). Confidence Intervals for the Mean of a Log-Normal Distribution. Journal of Statistics Education, 13(1). doi:10.1080/10691898.2005.11910638
Hothorn, T., & Lausen, B. (2003). On the Exact Distribution of Maximally Selected Rank Statistics. Computational Statistics & Data Analysis, 43(2), 121-137. doi:10.1016/S0167-9473(02)00225-6
Brown, L. D., Cai, T. T., & DasGupta, A. (2001). Interval Estimation for a Binomial Proportion. Statistical Science, 16(2), 101-133. doi:10.1214/ss/1009213286
Agresti, A., & Coull, B. A. (1998). Approximate is Better than "Exact" for Interval Estimation of Binomial Proportions. The American Statistician, 52(2), 119-126. doi:10.1080/00031305.1998.10480550
Newcombe, R. G. (1998). Two-Sided Confidence Intervals for the Single Proportion: Comparison of Seven Methods. Statistics in Medicine, 17(8), 857-872. doi:10.1002/sim.777
DiCiccio, T. J., & Efron, B. (1996). Bootstrap Confidence Intervals. Statistical Science, 11(3), 189-228. doi:10.1214/ss/1032280214
Frequently Asked Questions
What kinds of challenges is Sigma built for?
Sigma turns raw tabular data into a readable, statistically grounded binary tree. It is built to give a statistically robust, quick, fully automatic look at a dataset. It is not designed to compete with state-of-the-art but complex and time-consuming machine learning models on raw predictive performance. It is a good fit for reliable diagnostics on new datasets, one-off segmentation studies, and model drift investigations, and it helps examine the outputs of more complex models before relying on them.
How does Sigma differ from scikit-learn's decision trees?
scikit-learn's decision trees tend to be biased in their variable selection, require manual tuning of depth and minimum-samples thresholds to stay readable, and routinely grow into hairy, hard-to-interpret structures. Under the hood they pick splits by exhaustively optimizing a purity measure, which biases selection toward features with many possible split points and gives no statistically grounded stopping rule. Sigma uses permutation-based conditional-inference splits instead: at every node it tests each candidate variable for a real association with the response, picks the strongest, and stops the moment none is statistically significant. The tree finds its own depth automatically: no manual cutoff, no rule of thumb on samples per leaf. The single knob is the false-positive rate of each split test (the alpha level, 5% by default); raise it for a deeper exploratory tree, lower it for a stricter one. The result is a tree you can trust as a diagnostic.
Can I use Sigma for predictive modeling?
Yes, but I have something better. Fitting a Sigma tree gives you a robust first-approximation predictive model with very little effort: no tuning, no feature engineering, no encoding of categoricals, and a tree you can read end to end. It is a sound baseline you can deploy and trust. The tradeoff is subtlety for simplicity. A Sigma tree captures the dominant structure in the data but misses the finer interactions and gradients a richer model could exploit. When you need that accuracy without giving up interpretability, reach for Tau, my state-of-the-art engine for tabular data. It turns weak signals into outstanding predictions.
Does Sigma work with scikit-learn pipelines and Pandas DataFrames?
Yes. ClassificationTree, RegressionTree, and SurvivalTree implement the standard scikit-learn estimator API, so they slot into Pipeline, ColumnTransformer, GridSearchCV, cross-validation utilities, and model serialization without any custom glue. Pandas DataFrames are first-class inputs: pass them directly to fit and predict, declare categorical columns by name, and keep your column metadata throughout training and inference. NumPy arrays are of course also supported. Categorical features are handled natively (no upstream one-hot encoding needed), and sample weights are fully supported through all the statistical significance tests.
What confidence intervals does Sigma report on leaves?
Sigma supports the classic confidence intervals; the right one depends on the coverage guarantees you want. For classification: Clopper-Pearson, Jeffreys, and Wilson. For regression: Bayesian bootstrap, normal, Student-t, log-normal, gamma, Poisson, exponential, and beta. For survival outcomes: Brookmeyer-Crowley for the median survival time, log-log Greenwood for the survival probability at a fixed time, and Klein-Moeschberger integrated Greenwood for the restricted mean survival time (RMST). You pick the one that best fits your target's distribution.
Can Sigma handle very small or very large datasets?
On small datasets, the statistical stopping rule and the confidence intervals on leaves keep the tree honest about what it does and does not know. On larger datasets, fit time scales linearly with the number of observations and with the number of candidate variables, so Sigma's runtimes stay predictable as the data grows.
What Python versions and dependencies are required?
Python 3.10 and above, on every platform supported by NumPy and SciPy. Sigma depends only on NumPy, SciPy, and scikit-learn. Graphviz rendering is available as an optional extra for visualization, but it is not required to fit or use models.
How do I install Sigma?
A single pip install ars-sigma command installs
the library. There are
no compiled extensions to build, which
makes installation simple. Everything else (visualization
examples, deployment notes, source, and issue tracker) is on
the
GitHub repository.
What is Sigma's license, and where can I contribute?
Sigma is source-available: under the Sigma Source License you can freely read, use, modify, and redistribute the library, provided the license file and attribution travel with every copy. This is, however, not open source: ArsChitectura SAS retains all rights and may revoke the license at will. For broader rights and professional support, a commercial license is available on request through the contact page. Contributions, issues, and feedback are welcome on the GitHub repository.
How can I get support?
Reach out through the contact page. Your message goes directly to me.